
Research

Research Blog
"Can't start a fire without a spark"
Bruce Springsteen

Publications
-
Trans-regenerational RNAi Memory in Planarians
Prakash V. Cherian, Idit Aviram, Uri Weill, Tamar Shapira, Sarit Anava, Hila Gingold, Jochen C. Rink, Oded Rechavi, Omri Wurtzel bioRxiv 2026
-
Scavenger Cells Failure to Maintain Systemic RNA Homeostasis Causes Epigenetically Inherited Germline Tumors Itai Rieger, Yael Mor, Itamar Lev, Anat Nitzan, Charlotte Beata Kong, Sarit Anava, Hila Gingold, Ronen Zaidel-Bar, Oded Rechavi bioRxiv 2026
-
Cold and lithium delay forgetting of olfactory memories in Caenorhabditis elegans.
Landschaft-Berliner D, Goldstein K, Teichman G, Anava S, Gingold H, Salzberg Y, Rieger I, Levy N, Pechuk V, Setty H, Agarwal P, Sagi D, Cohen D, Nikelshparg E, Ben-Zvi A, Miranda-Vizuete A, Zaidel-Bar R, Oren-Suissa M, Rechavi O. Nat Neurosci. 2026.
-
S. Bracha, H. J. Johnson, N. A. Pranckevicius , F. Catto, A. E. Economides, S. Litvinov, K. Hassi, M. Tullio Rigoli, C. Cheroni, M. Bonfanti, A. Valenti , S. Stucchi, S. Attreya, P. D. Ross, D. Walsh, N. Malachi, H. Livne, R. Eshel, V. Krupalnik, D. Levin, S. Cobb, P. Koumoutsakos, N. Caporale, G. Testa, A. Aguzzi, A. A.Koshy, L. Sheiner, O. Rechavi. Engineering Toxoplasma gondii secretion systems for intracellular delivery of multiple large therapeutic proteins to neurons. Nature Microbiology, 2024.
-
I. Rieger, G. Weintraub, I. Lev, K. Goldstein, D. Bar-Zvi, S. Anava, H. Gingold, S. Shaham, O. Rechavi. Nucleus-Independent Transgenerational Small RNA Inheritance in C. elegans. Science Advances, 2023.
-
I. Toker, I. Lev, Y. Mor, Y. Gurevich, D. Fisher, L. Houri-Zeevi, O. Antonova, L. Hadany, S. Shaham, O. Rechavi. Transgenerational inheritance of sexual attractiveness via small RNAs enhances evolvability in C. elegans. Developmental Cell, 2022.
-
L. Houri-Ze’evi, G. Teichman, H. Gingold, O. Rechavi. “Stress Resets Ancestral Heritable Small RNA Responses". eLife, 2021.
-
R. Posner*, I.A. Toker*, O. Antonova, E. Star, S. Anava, E. Azmon, M. Hendricks, S. Bracha, H. Gingold, O. Rechavi. Neuronal Small RNAs Control Behavior Transgenerationally. Cell, 2019.
-
I. Lev*, I.A. Toker*, Y. Mor*, A. Nitzan, G. Weintraub, O. Antonova, O. Bhonkar, I. Ben Shushan, U. Seroussi, J.M. Claycomb, S. Anava, H. Gingold, R. Zaidel-Bar, O. Rechavi. Germ Granules Govern Small RNA Inheritance. Current Biology, 2019.
-
S. Anava, M. Neuhof, H. Gingold, O. Sagy, A. Munters, E. M.Svensson, E. Afshinnekoo, D. Danko, J. Foox, P. Shor, B. Riestra, D. Huchon, C.E. Mason, N. Mizrahi, M. Jakobsson, O. Rechavi. Illuminating Genetic, Mysteries of the Dead Sea Scrolls. Cell, 2020.
-
O.Rechavi, L. Houri-Ze'evi, S. Anava, W.S. Sho Goh, S.Y. Kerk, G.J. Hannon, O. Hobert. Starvation-induced transgenerational inheritance of small RNAs in C. elegans. Cell, 2014.

Research Tools

WorMachine
Machine Learning-Based Phenotypic Analysis Tool for Worms.
A three-step MATLAB-based image analysis software developed in the Rechavi Lab that allows automated identification of C. elegans worms, extraction of morphological features, and quantification of fluorescent signals.
Visit WorMachine
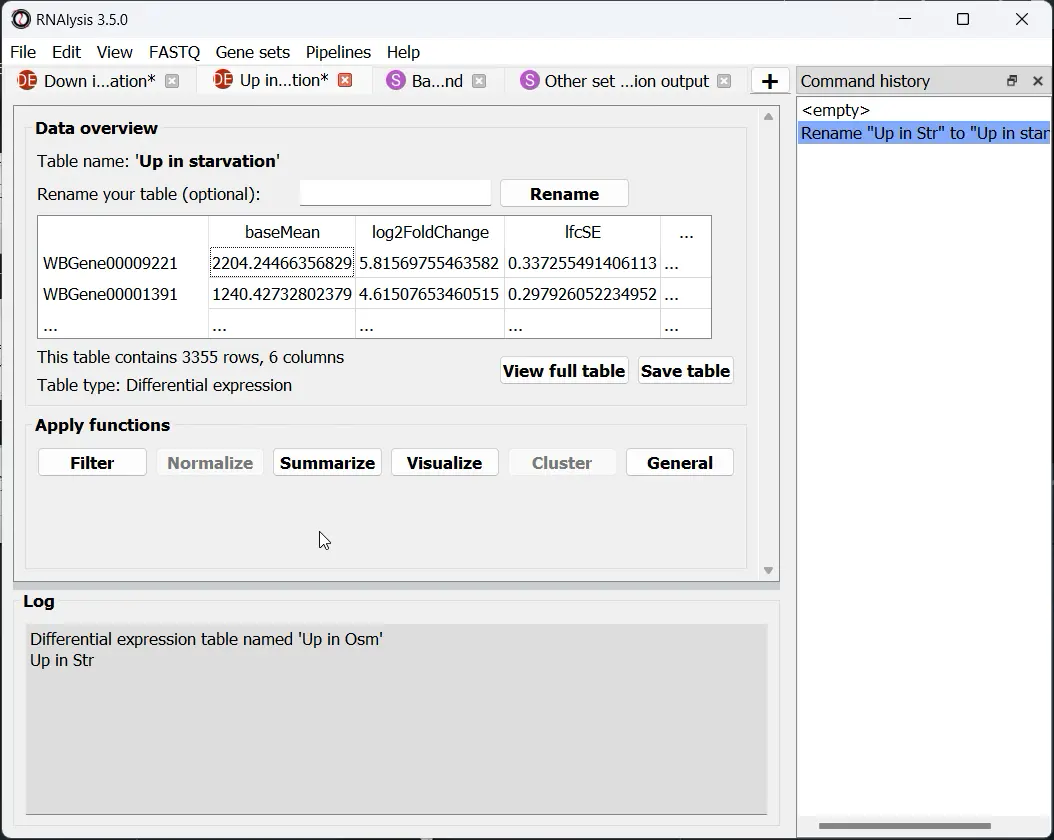
RNAlysis
RNAlysis is a modular Python-based analysis software for RNA sequencing data that was developed in the Oded Rechavi Lab.
RNAlysis allows users to build customized analysis pipelines suiting their specific research questions, going all the way from raw FASTQ files, through exploratory data analysis and data visualization, clustering analysis, and gene-set enrichment analysis. RNAlysis provides a friendly graphical user interface, allowing researchers to analyze data without writing code.
Visit RNAlysis

Funding






